Working with Senior Graphics Editor Jen Christiansen, Senior Editors Mark Fischetti and Clara Moskowitz, I have designed these Graphic Science visualizations for Scientific American.

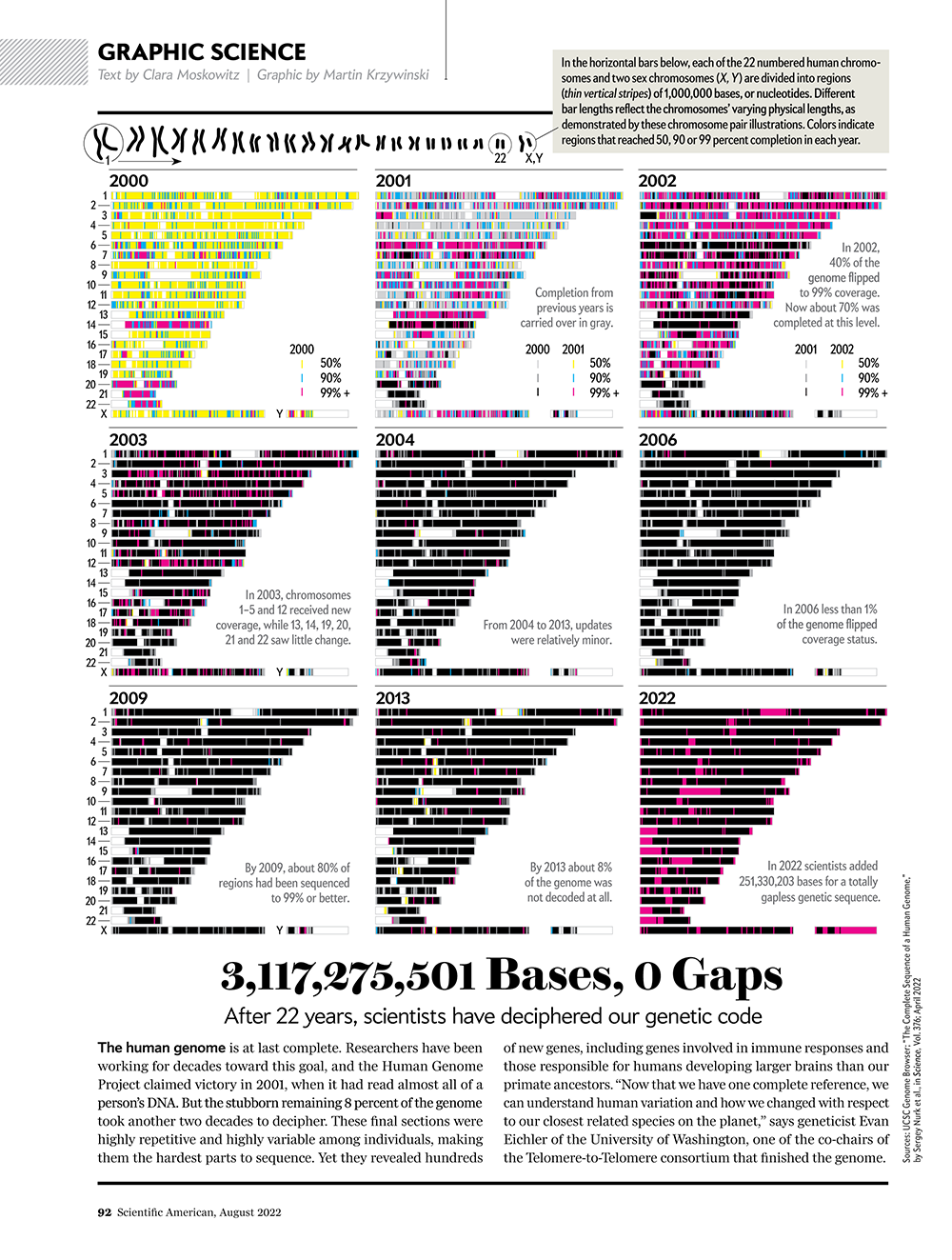
See how scientists put together the complete human genome
For the first time, researchers have sequenced all 3,117,275,501 bases of our genetic code.
Scientific American August 2022, volume 327 issue 2
text by Clara Moskowitz | graphic by Martin Krzywinski
source
assembly sequence from UCSC Genome Browser (assembly history), Nurk et al. The complete sequence of a human genome (2022) Science 376:44–53.
News
Filling in the Gaps by Laura Zahn, Most complete human genome yet reveals previously indecipherable DNA by Elizabeth Pennisi

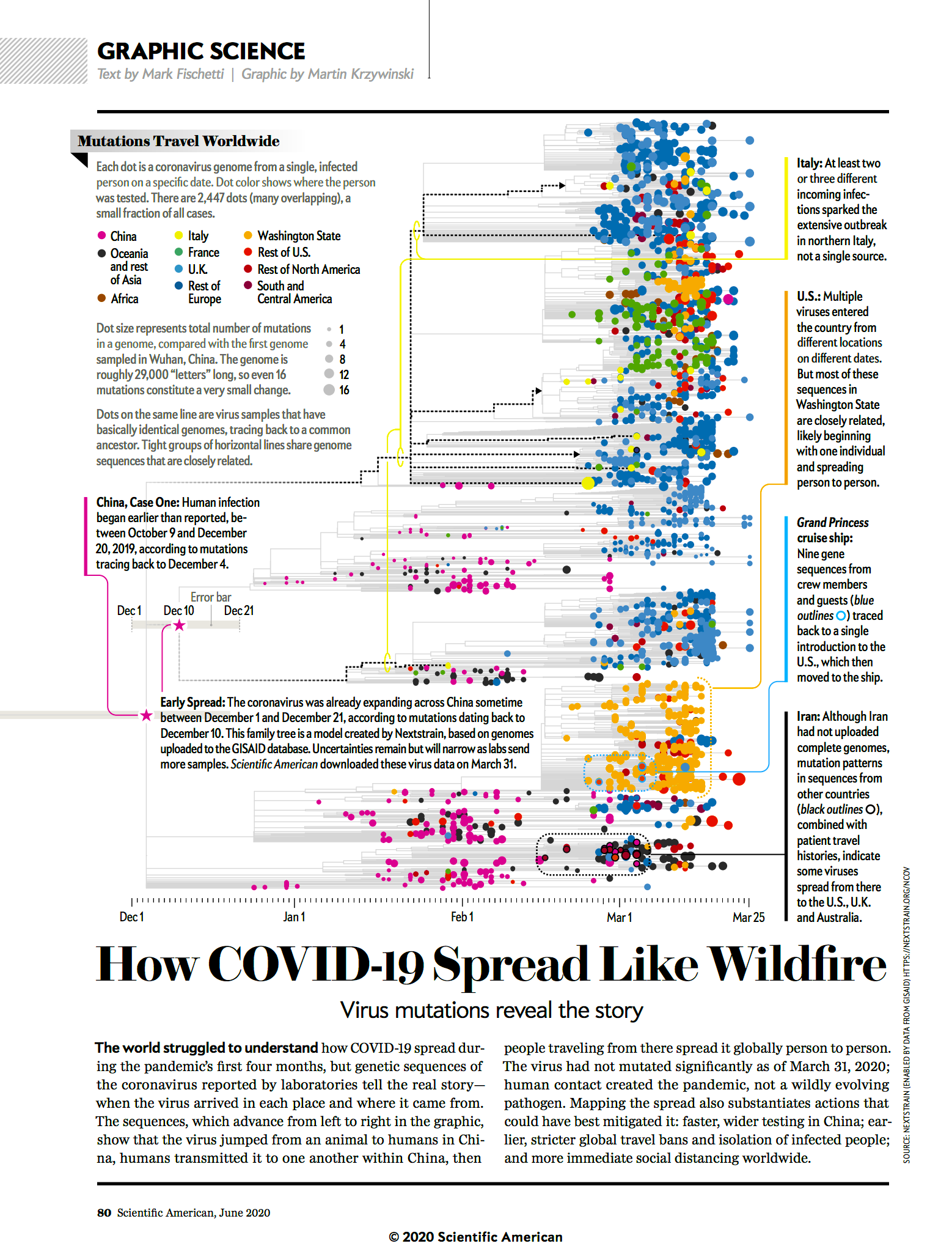
How COVID-19 spread like wildfire
Mutations in the SARS-Cov-2 virus reveal the story.
Scientific American June 2020, volume 322 issue 6
text by Mark Fischetti | graphic by Martin Krzywinski
Source
Interested in more COVID-19 and SARS-Cov-2 graphics? Check out my projects below.

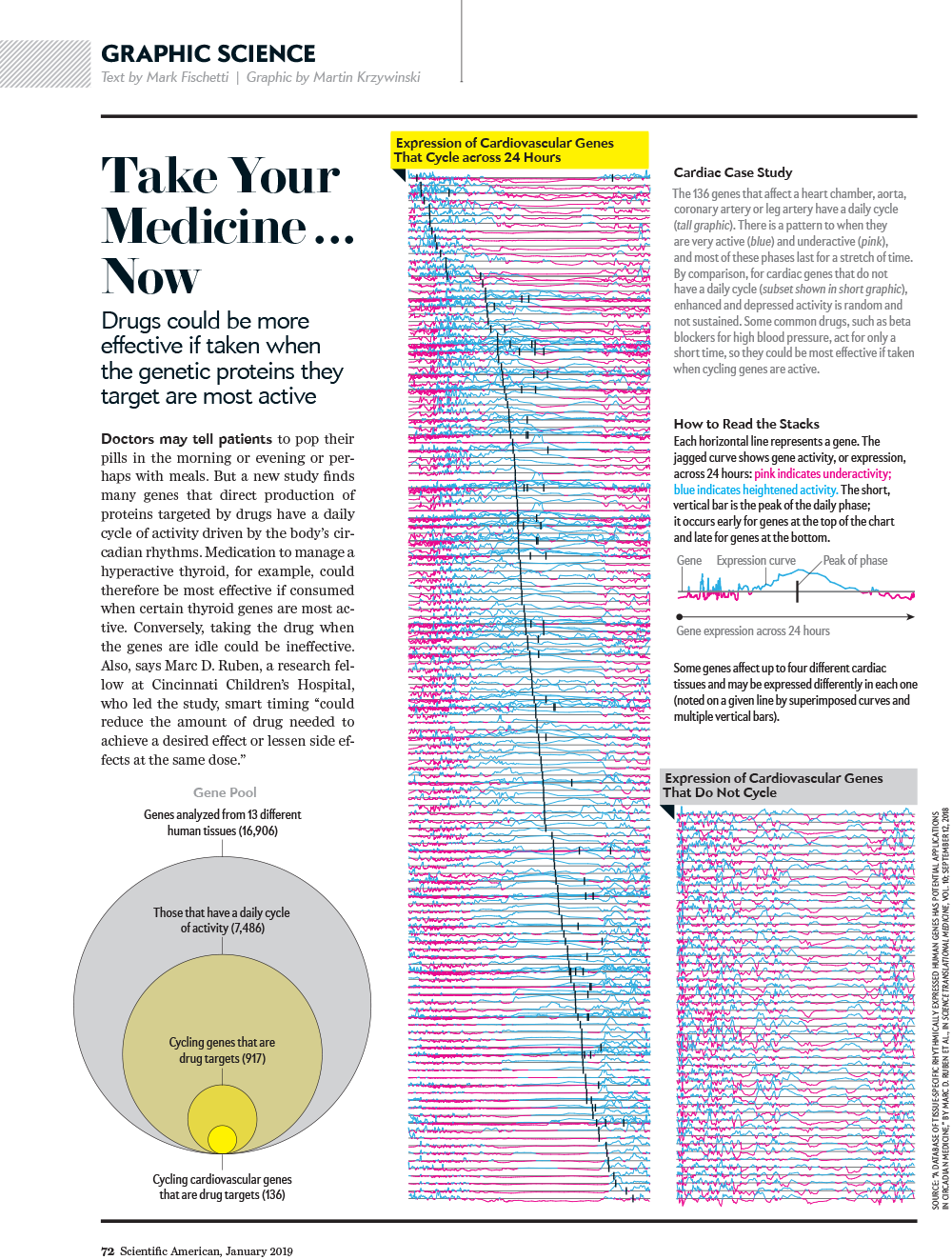
Take your medicine... now
Drugs could be more effective if taken when proteins they target are more active.
Scientific American January 2019, volume 320 issue 1
text by Mark Fischetti | graphic by Martin Krzywinski
A compelling overview figure of periodicity in a quantity. Before you even think about it, you already know what you're looking it.
Source
Ruben et al. A database of tissue-specific rhythmically expressed human genes has potential applications in circadian medicine Science Translational Medicine 10 Issue 458, eaat8806.

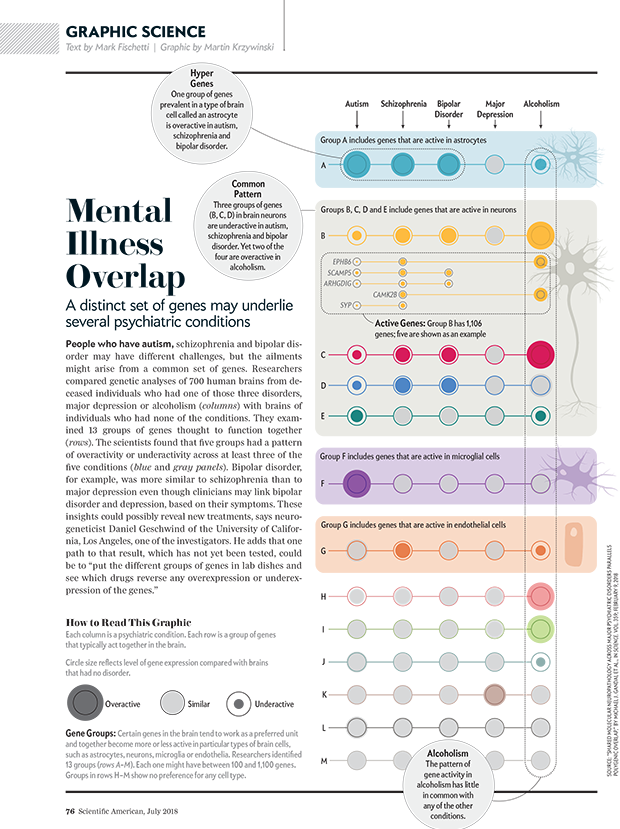
The same genes may underlie different psychiatric disorders
A distinct set of genes may underlie several psychiatric conditions.
Scientific American July 2018, volume 319 issue 1
text by Mark Fischetti | graphic by Martin Krzywinsk
The dataset is challenging: expression, correlation and network module membership of 11,000+ genes. Getting it onto one page was an exercise in restraint and calm.
source
Gandal M.J. et al. Shared Molecular Neuropathology Across Major Psychiatric Disorders Parallels Polygenic Overlap Science 359 693–697 (2018)

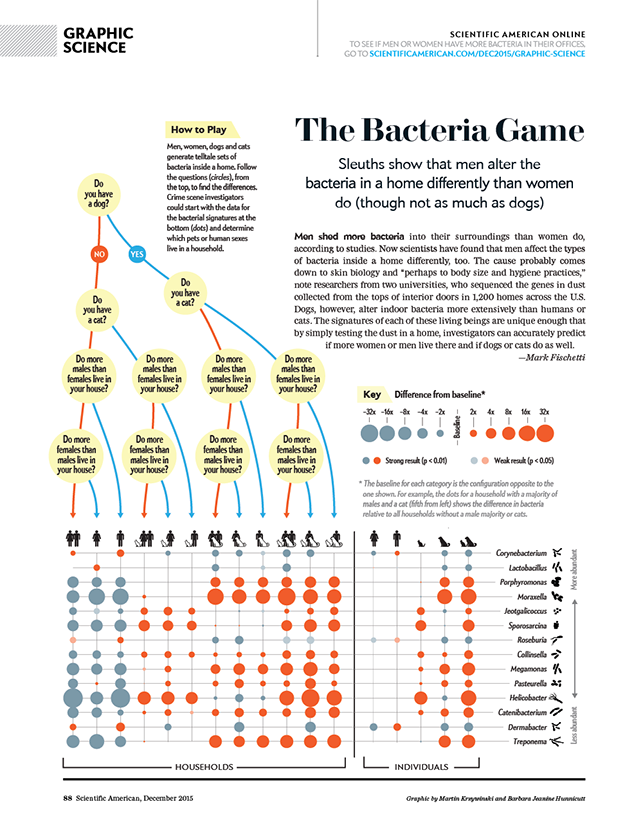
Men and women alter a home's bacteria differently
An analysis of dust reveals how the presence of men, women, dogs and cats affects the variety of bacteria in a household.
Scientific American December 2015, volume 313 issue 6
text by Mark Fischetti | graphic by Martin Krzywinski and Barbara Jeanine Hunnicutt
Catalogue of bacteria shapes by Barbara Jeanine Hunnicutt.
We explored differences in household dust bacteria based on the gender and pet status of the occupants.
We have also written about the making of the graphic, for those interested in how these things come together.
source
Barberan A et al. (2015) The ecology of microscopic life in household dust. Proc. R. Soc. B 282: 20151139.

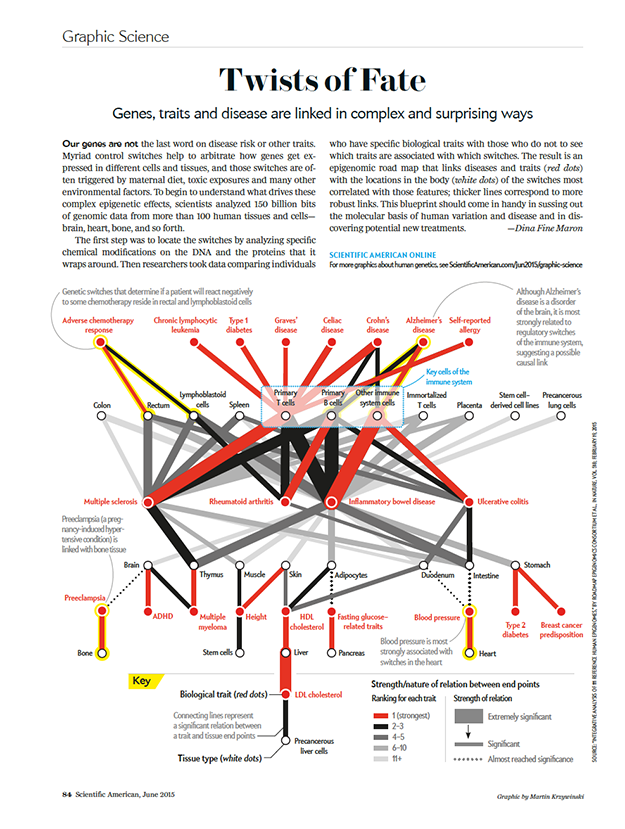
A roadmap to the "volume control" of genes
Genes, traits and disease are linked in complex and surprising ways.
Scientific American June 2015, volume 312 issue 6
text by Dina Fine Maron | graphic by Martin Krzywinski
Because sometimes only a network hairball will do.
source
Integrative analysis of 111 reference human epigenomes. (2015) Nature 518:317.

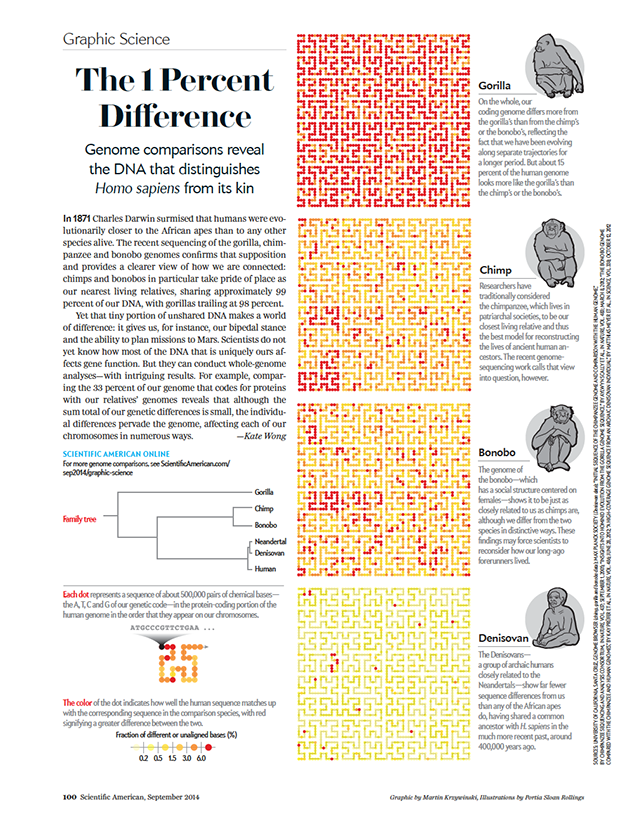
Tiny genetic differences between humans and other primates pervade the genome
Genome comparisons reveal the DNA that distinguishes Homo sapiens from its kin.
Scientific American September 2014, volume 311 issue 3
text by Kate Wong | illustrations by Portia Sloan Rollings | graphic by Martin Krzywinski
A Scientific American blog entry "A Monkey's Blueprint" accompanies this piece.
This design won a bronze award at Malofiej 23. For more information about Malofiej, see the SA Visual blog entry "There's No Infographic without Info (and other Lessons from Malofiej)".
A: Yes.
source

Nasa to send our human genome discs to the Moon
We'd like to say a ‘cosmic hello’: mathematics, culture, palaeontology, art and science, and ... human genomes.



Comparing classifier performance with baselines
All animals are equal, but some animals are more equal than others. —George Orwell
This month, we will illustrate the importance of establishing a baseline performance level.
Baselines are typically generated independently for each dataset using very simple models. Their role is to set the minimum level of acceptable performance and help with comparing relative improvements in performance of other models.

Unfortunately, baselines are often overlooked and, in the presence of a class imbalance5, must be established with care.
Megahed, F.M, Chen, Y-J., Jones-Farmer, A., Rigdon, S.E., Krzywinski, M. & Altman, N. (2024) Points of significance: Comparing classifier performance with baselines. Nat. Methods 20.
Happy 2024 π Day—
sunflowers ho!
Celebrate π Day (March 14th) and dig into the digit garden. Let's grow something.

How Analyzing Cosmic Nothing Might Explain Everything
Huge empty areas of the universe called voids could help solve the greatest mysteries in the cosmos.
My graphic accompanying How Analyzing Cosmic Nothing Might Explain Everything in the January 2024 issue of Scientific American depicts the entire Universe in a two-page spread — full of nothing.
The graphic uses the latest data from SDSS 12 and is an update to my Superclusters and Voids poster.
Michael Lemonick (editor) explains on the graphic:
“Regions of relatively empty space called cosmic voids are everywhere in the universe, and scientists believe studying their size, shape and spread across the cosmos could help them understand dark matter, dark energy and other big mysteries.
To use voids in this way, astronomers must map these regions in detail—a project that is just beginning.
Shown here are voids discovered by the Sloan Digital Sky Survey (SDSS), along with a selection of 16 previously named voids. Scientists expect voids to be evenly distributed throughout space—the lack of voids in some regions on the globe simply reflects SDSS’s sky coverage.”
voids
Sofia Contarini, Alice Pisani, Nico Hamaus, Federico Marulli Lauro Moscardini & Marco Baldi (2023) Cosmological Constraints from the BOSS DR12 Void Size Function Astrophysical Journal 953:46.
Nico Hamaus, Alice Pisani, Jin-Ah Choi, Guilhem Lavaux, Benjamin D. Wandelt & Jochen Weller (2020) Journal of Cosmology and Astroparticle Physics 2020:023.
Sloan Digital Sky Survey Data Release 12
Alan MacRobert (Sky & Telescope), Paulina Rowicka/Martin Krzywinski (revisions & Microscopium)
Hoffleit & Warren Jr. (1991) The Bright Star Catalog, 5th Revised Edition (Preliminary Version).
H0 = 67.4 km/(Mpc·s), Ωm = 0.315, Ωv = 0.685. Planck collaboration Planck 2018 results. VI. Cosmological parameters (2018).
constellation figures
stars
cosmology
Error in predictor variables
It is the mark of an educated mind to rest satisfied with the degree of precision that the nature of the subject admits and not to seek exactness where only an approximation is possible. —Aristotle
In regression, the predictors are (typically) assumed to have known values that are measured without error.
Practically, however, predictors are often measured with error. This has a profound (but predictable) effect on the estimates of relationships among variables – the so-called “error in variables” problem.

Error in measuring the predictors is often ignored. In this column, we discuss when ignoring this error is harmless and when it can lead to large bias that can leads us to miss important effects.
Altman, N. & Krzywinski, M. (2024) Points of significance: Error in predictor variables. Nat. Methods 20.
Background reading
Altman, N. & Krzywinski, M. (2015) Points of significance: Simple linear regression. Nat. Methods 12:999–1000.
Lever, J., Krzywinski, M. & Altman, N. (2016) Points of significance: Logistic regression. Nat. Methods 13:541–542 (2016).
Das, K., Krzywinski, M. & Altman, N. (2019) Points of significance: Quantile regression. Nat. Methods 16:451–452.
Convolutional neural networks
Nature uses only the longest threads to weave her patterns, so that each small piece of her fabric reveals the organization of the entire tapestry. – Richard Feynman
Following up on our Neural network primer column, this month we explore a different kind of network architecture: a convolutional network.
The convolutional network replaces the hidden layer of a fully connected network (FCN) with one or more filters (a kind of neuron that looks at the input within a narrow window).
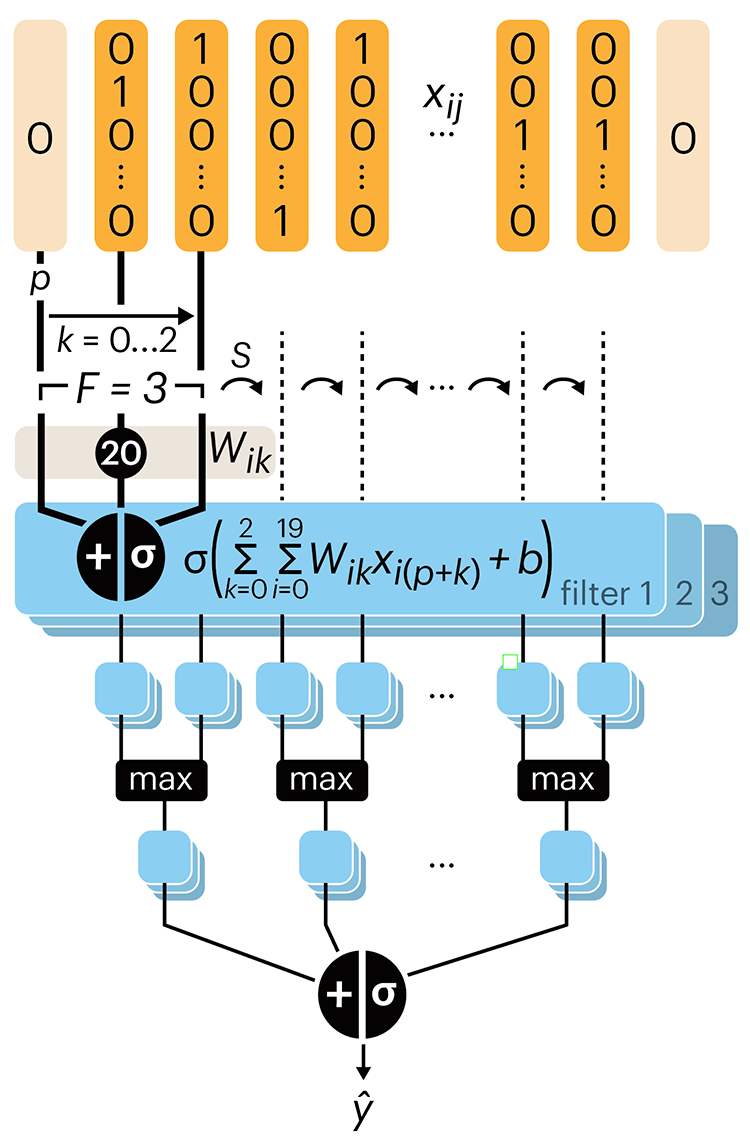
Even through convolutional networks have far fewer neurons that an FCN, they can perform substantially better for certain kinds of problems, such as sequence motif detection.
Derry, A., Krzywinski, M & Altman, N. (2023) Points of significance: Convolutional neural networks. Nature Methods 20:1269–1270.
Background reading
Derry, A., Krzywinski, M. & Altman, N. (2023) Points of significance: Neural network primer. Nature Methods 20:165–167.
Lever, J., Krzywinski, M. & Altman, N. (2016) Points of significance: Logistic regression. Nature Methods 13:541–542.
Neural network primer
Nature is often hidden, sometimes overcome, seldom extinguished. —Francis Bacon
In the first of a series of columns about neural networks, we introduce them with an intuitive approach that draws from our discussion about logistic regression.
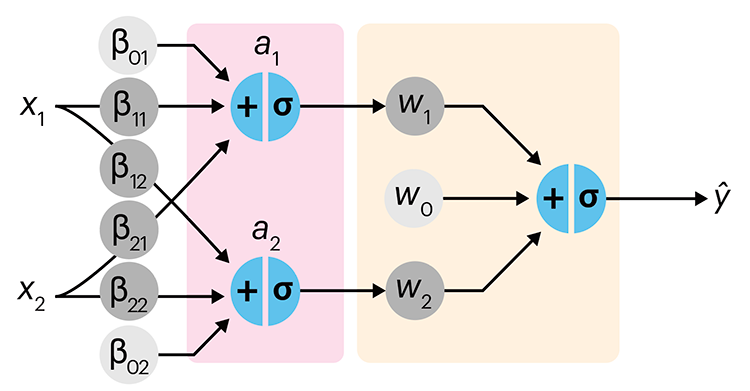
Simple neural networks are just a chain of linear regressions. And, although neural network models can get very complicated, their essence can be understood in terms of relatively basic principles.
We show how neural network components (neurons) can be arranged in the network and discuss the ideas of hidden layers. Using a simple data set we show how even a 3-neuron neural network can already model relatively complicated data patterns.
Derry, A., Krzywinski, M & Altman, N. (2023) Points of significance: Neural network primer. Nature Methods 20:165–167.
Background reading
Lever, J., Krzywinski, M. & Altman, N. (2016) Points of significance: Logistic regression. Nature Methods 13:541–542.
Cell Genomics cover
Our cover on the 11 January 2023 Cell Genomics issue depicts the process of determining the parent-of-origin using differential methylation of alleles at imprinted regions (iDMRs) is imagined as a circuit.
Designed in collaboration with with Carlos Urzua.

Akbari, V. et al. Parent-of-origin detection and chromosome-scale haplotyping using long-read DNA methylation sequencing and Strand-seq (2023) Cell Genomics 3(1).
Browse my gallery of cover designs.