Schemaball
Circular visualization of database schemas
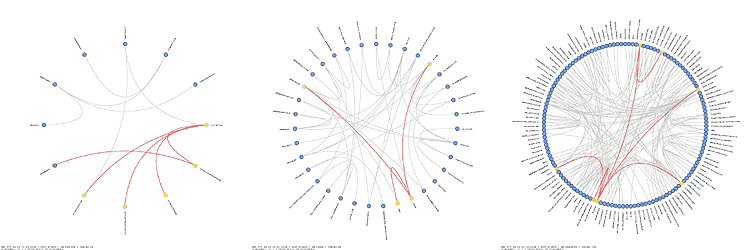
Schemaball was published in SysAdmin Magazine (Krzywinski, M. Schemaball: A New Spin on Database Visualization (2004) Sysadmin Magazine Vol 13 Issue 08). Who cites Schemaball?
These tutorials illustrate different features of Schemaball. Tutorials are divided into sections. Each section covers a particular aspect of a feature or set of features. Each section has its own documentation, configuration file and schema ball image.
tutorial 1
Fetching, parsing and viewing an SQL schema from a remote server.
NAME
sbtut_01 - fetching, parsing and viewing an SQL schema from a remote server
SYNOPSIS
> mysql -u anonymous -h kaka.sanger.ac.uk -e "show databases" > databases.txt > grep homo_sapiens_core databases.txt homo_sapiens_core_19_34b homo_sapiens_core_20_34c homo_sapiens_core_21_34d > mysqldump -u anonymous -h kaka.sanger.ac.uk -d -n homo_sapiens_core_21_34d > hs.sql
PURPOSE
To introduce Schemaball, we'll download an SQL schema dump for a Ensembl human genome database. We'll create a simple configuration file, parse the database and create a schema diagram.
TUTORIAL
For this tutorial, the Ensembl database (www.ensembl.org) for the human genome will be used as an example. Ensembl is an open-source system for storing, analyzing and visualizing genomic data for any organism. It is a very popular public resource.
The diagram created in this tutorial will not contain any constraint lines. Tutorial 2 will show you how to parse constraint relationships from the schema and add them to the diagram.
Obtaining an SQL dump
Ensembl provides a public MySQL server, making it possible to query the database directly.
host: kaka.sanger.ac.uk user: anonymous pass: <NO PASSWORD>
The Ensembl server contains a large number of databases. Data for each species stored in several databases. See www.ensembl.org/Docs for details. To list the databases at Ensembl
> mysql -u anonymous -h kaka.sanger.ac.uk -e "show databases"
Let's look at the tables with data for the human (homo sapiens) genome. At any point, there may be multiple versions of the database available. For example, at the time of writing, the ``core'' human database was available in three versions
> homo_sapiens_core_19_34b > homo_sapiens_core_20_34c > homo_sapiens_core_21_34d
Version 21 is comprised of these databases
> homo_sapiens_core_21_34d > homo_sapiens_disease_21_34d > homo_sapiens_est_21_34d > homo_sapiens_estgene_21_34d > homo_sapiens_haplotype_21_34d > homo_sapiens_lite_21_34d > homo_sapiens_snp_21_34d > homo_sapiens_vega_21_34d
Let's fetch the schema for the _core_ database.
> mysqldump -u anonymous -h kaka.sanger.ac.uk -d -n homo_sapiens_core_21_34d
It's important to use -d, so that only the table structure is downloaded and not the data, and -n, to avoid the database creation statement which can confuse the SQL::Translator module used by Schemaball.
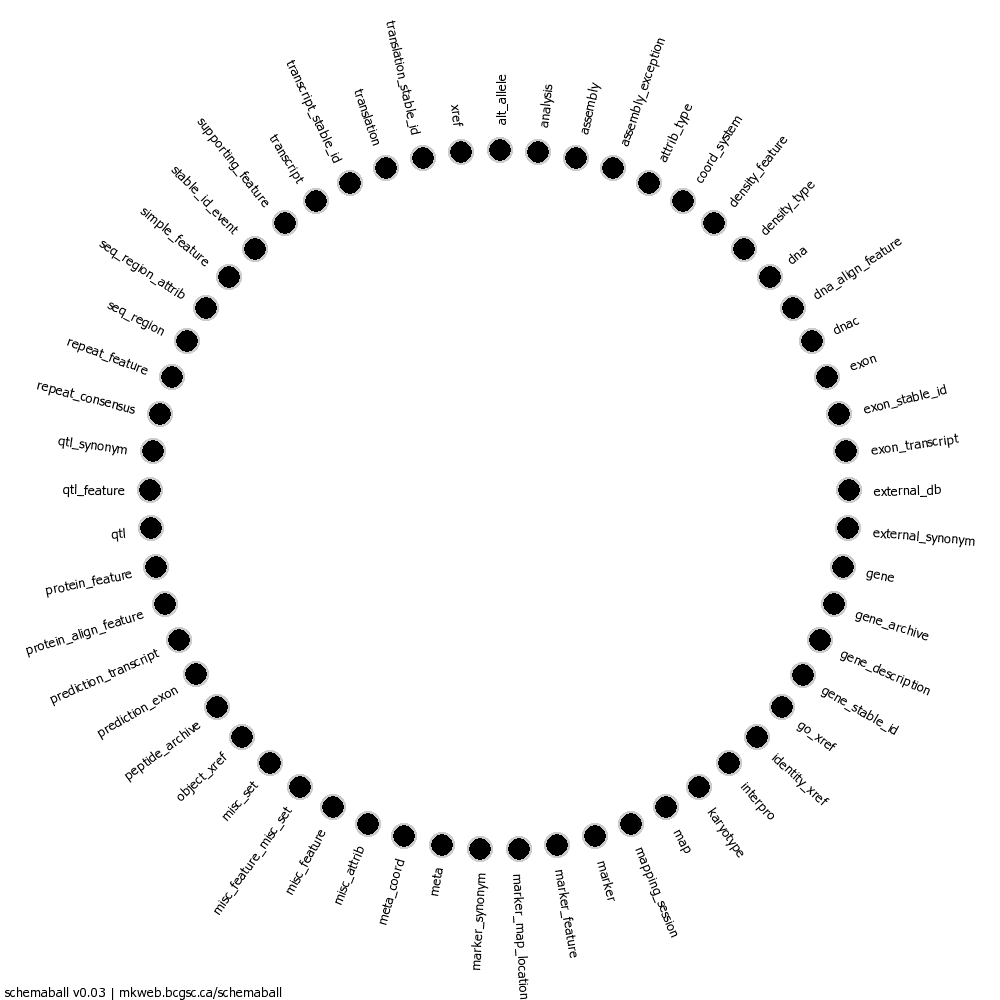
The Homo Sapiens Core SQL Dump
The SQL schema dump consists of all the necessary CREATE TABLE directives needed to reconstruct the structure of the database. One such table is
CREATE TABLE karyotype (
karyotype_id int(10) unsigned NOT NULL auto_increment,
seq_region_id int(10) unsigned NOT NULL default '0',
seq_region_start int(10) NOT NULL default '0',
seq_region_end int(10) NOT NULL default '0',
band varchar(40) NOT NULL default '',
stain varchar(40) NOT NULL default '',
PRIMARY KEY (karyotype_id),
KEY region_band_idx (seq_region_id,band)
) TYPE=MyISAM;
Ensembl uses MyISAM tables, which do not support foreign keys in MySQL 4.0. By looking at the schema, you'll see that the primary key usually has the format TABLENAME_id and that foreign keys have the format TABLENAME_FIELD. For example, the karotype table uses karotype_id as its primary key and points to the seq_region table using the field seq_region_id. Because the schema is does not store these foreign keys using constraint options, you'll need to tell Schemaball how to figure out which fields are foreign keys in order to work out what the constraint relationships are. This process is addressed in tutorial 2 (sbtut_02).
Configuring Schemaball
Schemaball supports many configuration options. Many of them are optional and a small configuration file is required to get the job done. The configuration file for this tutorial, sbtut_01.conf, specifies the schema file in a <db> block and image parameters in an <image> block.
<db> sqlfile = tutorial/01/hs.sql </db>
The image parameters direct where and how the table glyphs should be constructed. Since we don't have any foreign key relationships parsed, only table options need to be included.
<image>
file = /home/httpd/htdocs/worldatwar/schema.png
size = 1000
glyphradius = 0.7
background = ffffff
foreground = 000000
<elements>
<label>
font = tahoma.ttf
size = 9
</label>
<table>
size = 25
fill = 000000
stroke = 2
outline = cccccc
</table>
</elements>
</image>
Parsing the Schema
To parse the schema and produce the schema diagram,
> schemaball -conf tutorial/01/sbtut_01.conf -debug > parsed.txt
The diagram is included as sbtut_01.png.
Schemaball generates useful information about the schema when the -debug flag is used.
- table list
The structure of the schema, which includes the list of tables and constraints, is reported in ``schemadump'' lines.
> grep schemadump parsed.txt debug schemadump table alt_allele debug schemadump table analysis debug schemadump table assembly ...
At this point, these lines do not provide very useful information. First, we don't know anything about constraints (yet) and the table names can be easily parsed from the hs.sql file.
- element coordinates
The coordinates of each element drawn by schemaball are reported in ``ballelem'' lines.
> grep ballelem parsed.txt ballelem type text xy 506 125 rt 375 0.01733 rot 1.57080 size 25 label alt_allele ballelem type circle xy 500 150 rt 350 0.00000 rot 0.00000 size 25 label alt_allele
These lines can be parsed to generate SVG images.
HISTORY
BUGS
Please report errors and suggestions to martink@bcgsc.ca
AUTHOR
Martin Krzywinski, mkweb.bcgsc.ca, martink@bcgsc.ca
$Id: schemaball,v 1.5 2003/10/27 20:25:11 martink Exp $
CONTACT
Martin Krzywinski Genome Sciences Centre Vancouver BC Canada www.bcgsc.ca mkweb.bcgsc.ca martink@bcgsc.ca
configuration
################################################################ # # Schemaball configuration file # # Tutorial 01 # # sbtut_01_01.conf - fetching and parsing SQL schemas from a remote server # # The remote server used here is the Ensembl server (www.ensembl.org). # The database is the Homo sapiens core database. # <db> sqlfile = tutorial/01/hs.sql </db> <image> file = /home/httpd/htdocs/worldatwar/schema.png size = 1000 glyphradius = 0.7 image_element_dump = yes background = ffffff foreground = 000000 <elements> <label> font = tahoma.ttf size = 10 </label> <table> size = 25 fill = 000000 stroke = 2 outline = cccccc </table> </elements> </image>

Nasa to send our human genome discs to the Moon
We'd like to say a ‘cosmic hello’: mathematics, culture, palaeontology, art and science, and ... human genomes.



Comparing classifier performance with baselines
All animals are equal, but some animals are more equal than others. —George Orwell
This month, we will illustrate the importance of establishing a baseline performance level.
Baselines are typically generated independently for each dataset using very simple models. Their role is to set the minimum level of acceptable performance and help with comparing relative improvements in performance of other models.

Unfortunately, baselines are often overlooked and, in the presence of a class imbalance5, must be established with care.
Megahed, F.M, Chen, Y-J., Jones-Farmer, A., Rigdon, S.E., Krzywinski, M. & Altman, N. (2024) Points of significance: Comparing classifier performance with baselines. Nat. Methods 20.
Happy 2024 π Day—
sunflowers ho!
Celebrate π Day (March 14th) and dig into the digit garden. Let's grow something.

How Analyzing Cosmic Nothing Might Explain Everything
Huge empty areas of the universe called voids could help solve the greatest mysteries in the cosmos.
My graphic accompanying How Analyzing Cosmic Nothing Might Explain Everything in the January 2024 issue of Scientific American depicts the entire Universe in a two-page spread — full of nothing.
The graphic uses the latest data from SDSS 12 and is an update to my Superclusters and Voids poster.
Michael Lemonick (editor) explains on the graphic:
“Regions of relatively empty space called cosmic voids are everywhere in the universe, and scientists believe studying their size, shape and spread across the cosmos could help them understand dark matter, dark energy and other big mysteries.
To use voids in this way, astronomers must map these regions in detail—a project that is just beginning.
Shown here are voids discovered by the Sloan Digital Sky Survey (SDSS), along with a selection of 16 previously named voids. Scientists expect voids to be evenly distributed throughout space—the lack of voids in some regions on the globe simply reflects SDSS’s sky coverage.”
voids
Sofia Contarini, Alice Pisani, Nico Hamaus, Federico Marulli Lauro Moscardini & Marco Baldi (2023) Cosmological Constraints from the BOSS DR12 Void Size Function Astrophysical Journal 953:46.
Nico Hamaus, Alice Pisani, Jin-Ah Choi, Guilhem Lavaux, Benjamin D. Wandelt & Jochen Weller (2020) Journal of Cosmology and Astroparticle Physics 2020:023.
Sloan Digital Sky Survey Data Release 12
Alan MacRobert (Sky & Telescope), Paulina Rowicka/Martin Krzywinski (revisions & Microscopium)
Hoffleit & Warren Jr. (1991) The Bright Star Catalog, 5th Revised Edition (Preliminary Version).
H0 = 67.4 km/(Mpc·s), Ωm = 0.315, Ωv = 0.685. Planck collaboration Planck 2018 results. VI. Cosmological parameters (2018).
constellation figures
stars
cosmology
Error in predictor variables
It is the mark of an educated mind to rest satisfied with the degree of precision that the nature of the subject admits and not to seek exactness where only an approximation is possible. —Aristotle
In regression, the predictors are (typically) assumed to have known values that are measured without error.
Practically, however, predictors are often measured with error. This has a profound (but predictable) effect on the estimates of relationships among variables – the so-called “error in variables” problem.

Error in measuring the predictors is often ignored. In this column, we discuss when ignoring this error is harmless and when it can lead to large bias that can leads us to miss important effects.
Altman, N. & Krzywinski, M. (2024) Points of significance: Error in predictor variables. Nat. Methods 20.
Background reading
Altman, N. & Krzywinski, M. (2015) Points of significance: Simple linear regression. Nat. Methods 12:999–1000.
Lever, J., Krzywinski, M. & Altman, N. (2016) Points of significance: Logistic regression. Nat. Methods 13:541–542 (2016).
Das, K., Krzywinski, M. & Altman, N. (2019) Points of significance: Quantile regression. Nat. Methods 16:451–452.
