Schemaball
Circular visualization of database schemas
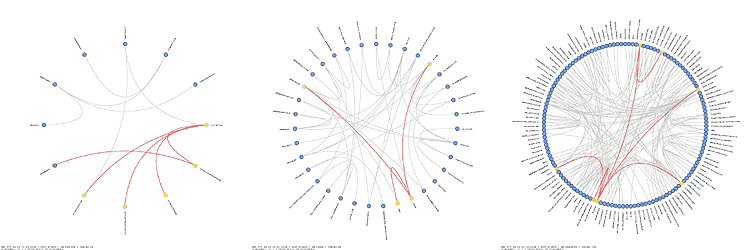
Schemaball was published in SysAdmin Magazine (Krzywinski, M. Schemaball: A New Spin on Database Visualization (2004) Sysadmin Magazine Vol 13 Issue 08). Who cites Schemaball?
overview
Schemaball is a Perl script which uses GD to generate static, circularly formatted SQL database schema views. Schemaball is well-suited for use in publications, online or print, presentations or schema development. Schemaball is suitable for visualizing schemas of all sizes.
features
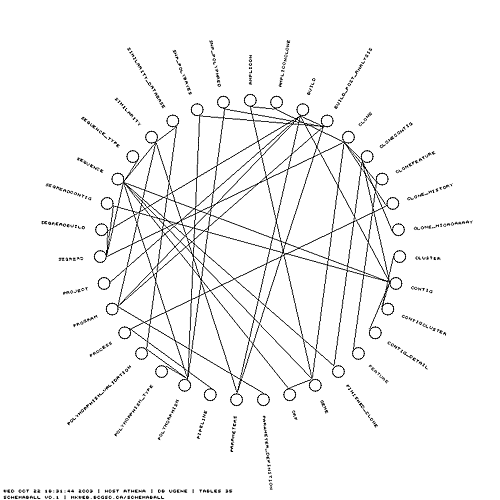
To illustrate the features of schemaball, I'll use our ugene MySQL database (middle schema in the above figure). In the schema view below, the tables can be seen organized around a circle in alphabetical order. Tables which are linked by foreign key relationships — I'll get to how this is done in MySQL in a moment — are linked using bezier curves. In the view below, neighbours and next-neighbours of the Clone table are highlighted.
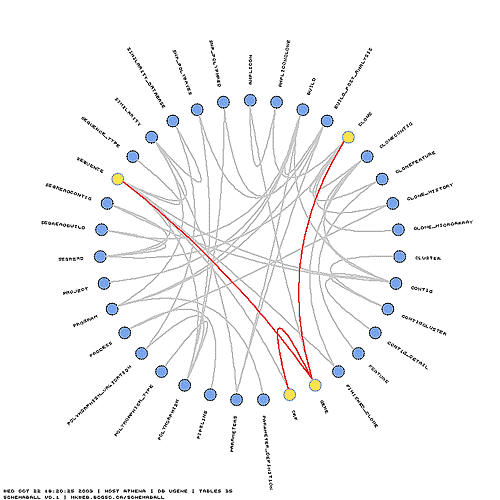
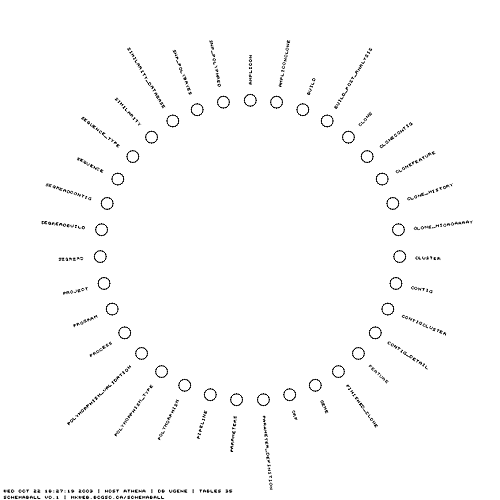
The simplest schema ball is one in which the table glyphs are shown without links. I find this view to be useful as a design template. I can print a few copies out and draw possible table relationships as I work on a schema.
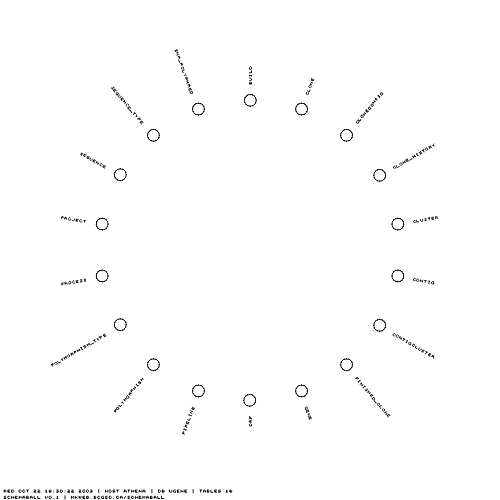
Tables can be hidden from the schema ball using regular expressions that are applied to the table names. In this case I've removed all tables which have the letter "a" in them.
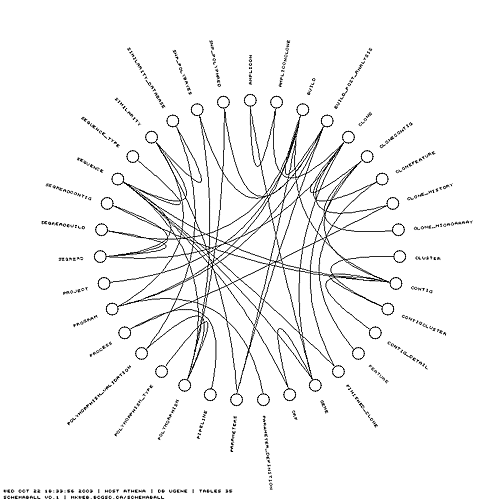
One of the main features of schemaball is the ability to visualize links between tables. Schemaball can parse these links from the schema structure itself (CONSTRAINT table options), using field names (if you've named your foreign keys using some convention), or from a file listing the table pairs.

The curvature of the Bezier lines can be adjusted using a parameter which controls the distance from the center of the schema ball of the middle point on the Bezier curve.
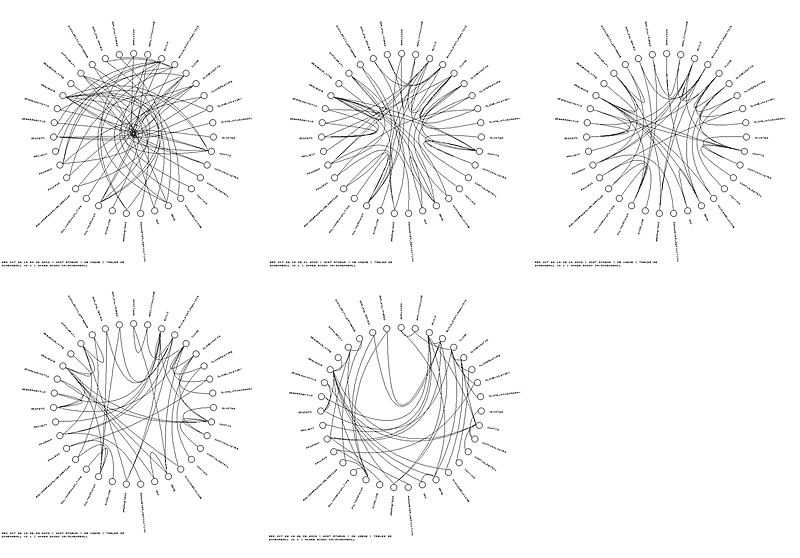
Schemaball is very flexible. You can turn off the table glyphs and labels and have your schema turned into Bezier art.

If you are paranoid about your intellectual property but would like to show the complexity of your schema to impress your competitors, you can anonymize the labels.
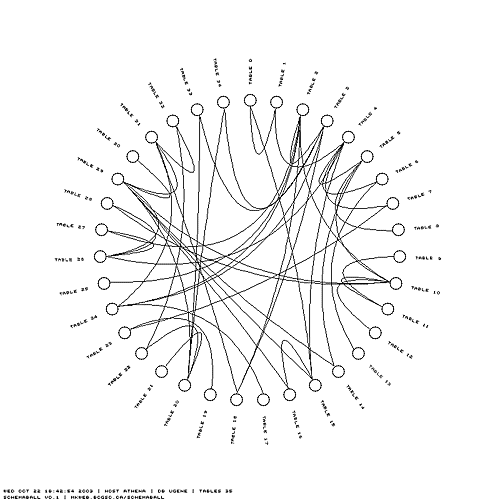

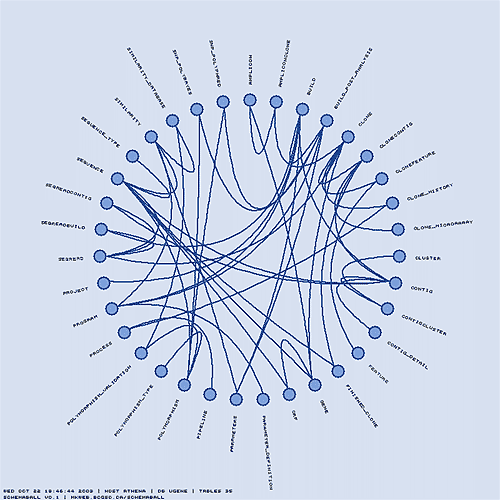
You can specify the colour characteristics of the features in the schema ball. Let's switch to a soothing blue theme. The table glyphs can be stroked and the link lines made thicker.
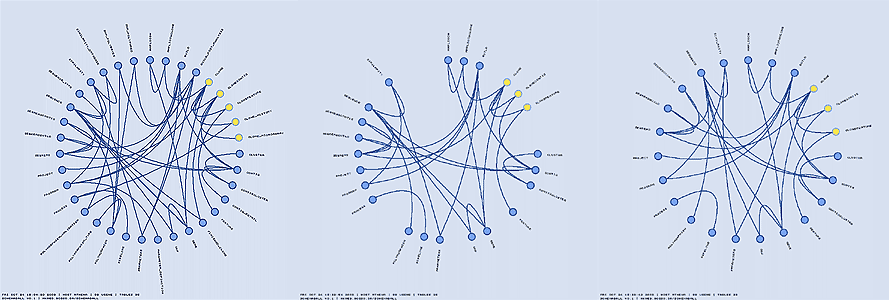
Tables can be highlighted using regular expressions, which are applied to the table names. Below, I show how three different table groups can be highlighted.
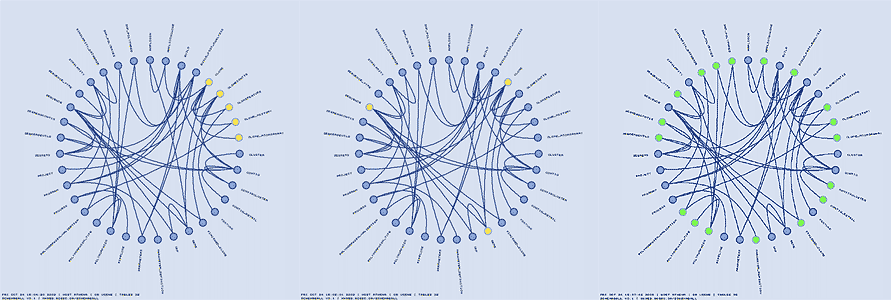

Tables can be hidden from the schema ball. This feature is useful if you have a lot of tables and would like to focus on the relationships between a subset of tables. Hiding is implemented in two ways: making tables in the schema ball invisible, but retaining a gap in the schema ball where the invisible table glyph is, or removing the table from the schema ball altogether and rearranging the other tables to fill the ball. Hiding is controlled using regular expression, in the same way as highlighting.
If you hide tables by making them invisible, you can choose to still have links to these tables kept in the schema ball. I don't know why you'd want to do this, but you can.
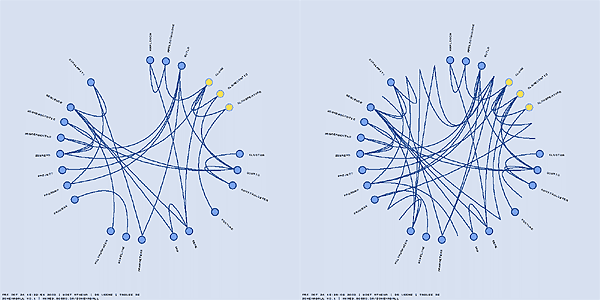
In addition to hiding and highlighting tables, you can also hide and highlight links. The hiding and highlighting process is controlled by regular expressions, like for tables. A link that joins two tables TABLE1 and TABLE2 together is named TABLE1___TABLE2 and the regular expressions controlling link visibility are applied to this compound name. For example, "^TABLE1___" selects all links which point from TABLE1 and "___TABLE2$" selects all links which point to TABLE2.
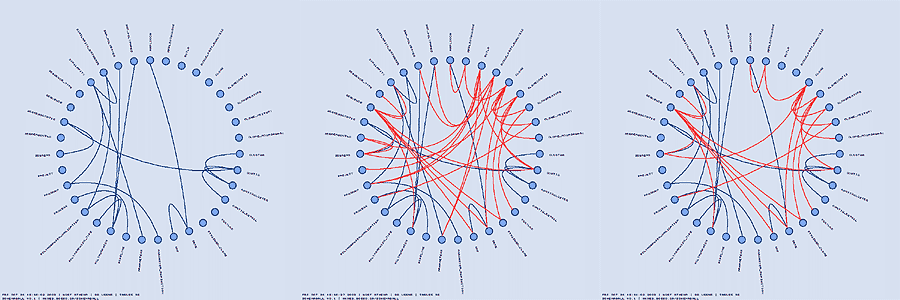

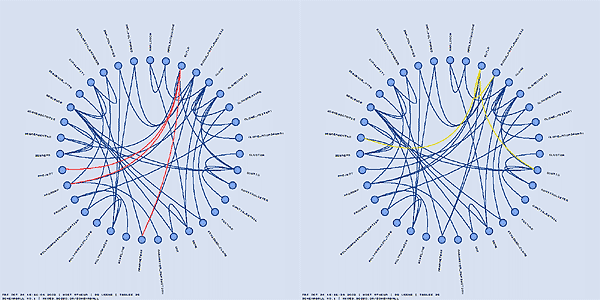
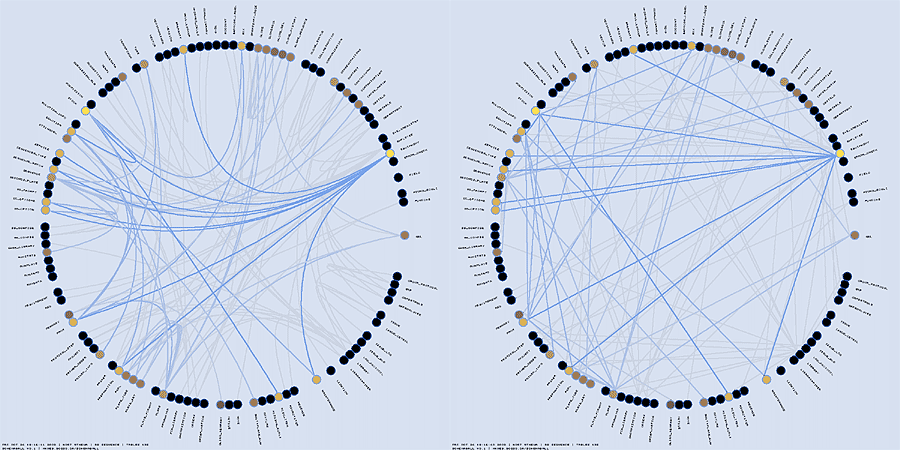
Tracing the table dependency through foreign key relationships can be tedious in a large schema. Schemaball supports a chain hilighting scheme which follows links from highlighted tables and highlights connected tables. You can adjust the number of iterations of this scheme to highlight linked neighbours of a table to varying degrees of separation. A number of different parameters controls how the highlighting is inherited.
The "highlight_by_link" property is used to highlight tables which connect to highlighted links. This is useful if you would like to highlight all tables which, for example, are referenced by a specific table.
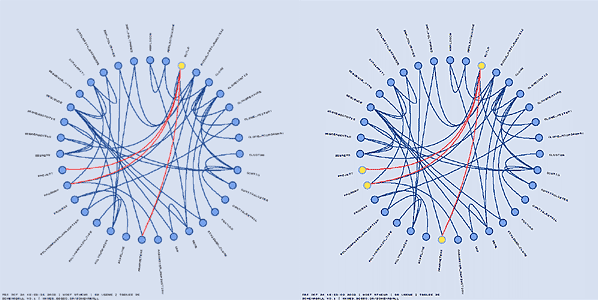
The highlight_by_iterations specifies the number of cycles of highlight inheritance that Schemaball should follow. You can follow links in the forward or reverse direction, or both.
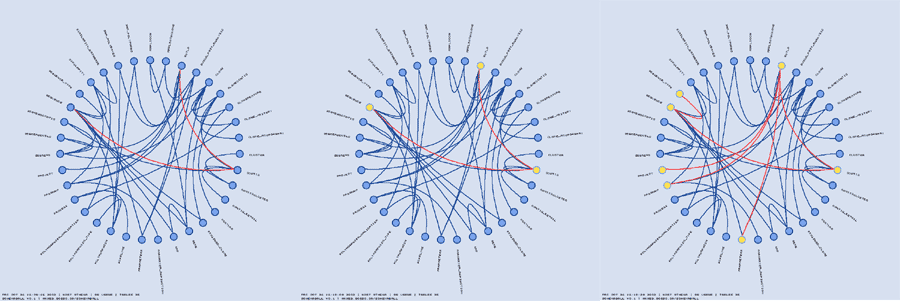
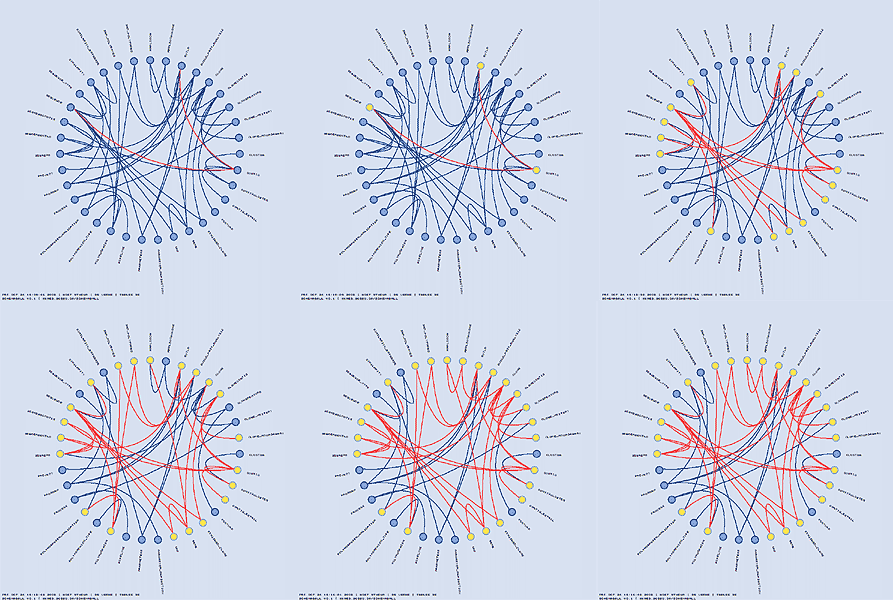
If a large value for highlight_by_iterations is used and the schema is large, you can wind up with many highlighted elements. In order to retain information about the inheritance depth of a highlighted element, the "fade_factor_table" and "fade_factor_link" parameters are used. When these parameters are used, the highlight colour is progressively diluted with each iteration.
Nasa to send our human genome discs to the Moon
We'd like to say a ‘cosmic hello’: mathematics, culture, palaeontology, art and science, and ... human genomes.



Comparing classifier performance with baselines
All animals are equal, but some animals are more equal than others. —George Orwell
This month, we will illustrate the importance of establishing a baseline performance level.
Baselines are typically generated independently for each dataset using very simple models. Their role is to set the minimum level of acceptable performance and help with comparing relative improvements in performance of other models.

Unfortunately, baselines are often overlooked and, in the presence of a class imbalance5, must be established with care.
Megahed, F.M, Chen, Y-J., Jones-Farmer, A., Rigdon, S.E., Krzywinski, M. & Altman, N. (2024) Points of significance: Comparing classifier performance with baselines. Nat. Methods 20.
Happy 2024 π Day—
sunflowers ho!
Celebrate π Day (March 14th) and dig into the digit garden. Let's grow something.

How Analyzing Cosmic Nothing Might Explain Everything
Huge empty areas of the universe called voids could help solve the greatest mysteries in the cosmos.
My graphic accompanying How Analyzing Cosmic Nothing Might Explain Everything in the January 2024 issue of Scientific American depicts the entire Universe in a two-page spread — full of nothing.
The graphic uses the latest data from SDSS 12 and is an update to my Superclusters and Voids poster.
Michael Lemonick (editor) explains on the graphic:
“Regions of relatively empty space called cosmic voids are everywhere in the universe, and scientists believe studying their size, shape and spread across the cosmos could help them understand dark matter, dark energy and other big mysteries.
To use voids in this way, astronomers must map these regions in detail—a project that is just beginning.
Shown here are voids discovered by the Sloan Digital Sky Survey (SDSS), along with a selection of 16 previously named voids. Scientists expect voids to be evenly distributed throughout space—the lack of voids in some regions on the globe simply reflects SDSS’s sky coverage.”
voids
Sofia Contarini, Alice Pisani, Nico Hamaus, Federico Marulli Lauro Moscardini & Marco Baldi (2023) Cosmological Constraints from the BOSS DR12 Void Size Function Astrophysical Journal 953:46.
Nico Hamaus, Alice Pisani, Jin-Ah Choi, Guilhem Lavaux, Benjamin D. Wandelt & Jochen Weller (2020) Journal of Cosmology and Astroparticle Physics 2020:023.
Sloan Digital Sky Survey Data Release 12
Alan MacRobert (Sky & Telescope), Paulina Rowicka/Martin Krzywinski (revisions & Microscopium)
Hoffleit & Warren Jr. (1991) The Bright Star Catalog, 5th Revised Edition (Preliminary Version).
H0 = 67.4 km/(Mpc·s), Ωm = 0.315, Ωv = 0.685. Planck collaboration Planck 2018 results. VI. Cosmological parameters (2018).
constellation figures
stars
cosmology
Error in predictor variables
It is the mark of an educated mind to rest satisfied with the degree of precision that the nature of the subject admits and not to seek exactness where only an approximation is possible. —Aristotle
In regression, the predictors are (typically) assumed to have known values that are measured without error.
Practically, however, predictors are often measured with error. This has a profound (but predictable) effect on the estimates of relationships among variables – the so-called “error in variables” problem.

Error in measuring the predictors is often ignored. In this column, we discuss when ignoring this error is harmless and when it can lead to large bias that can leads us to miss important effects.
Altman, N. & Krzywinski, M. (2024) Points of significance: Error in predictor variables. Nat. Methods 20.
Background reading
Altman, N. & Krzywinski, M. (2015) Points of significance: Simple linear regression. Nat. Methods 12:999–1000.
Lever, J., Krzywinski, M. & Altman, N. (2016) Points of significance: Logistic regression. Nat. Methods 13:541–542 (2016).
Das, K., Krzywinski, M. & Altman, N. (2019) Points of significance: Quantile regression. Nat. Methods 16:451–452.
