Nature Methods: Points of Significance

Access all columns for free at Statistics for Biologists Nature Collection.
A Statistics Primer and Best Practices
The Points of Significance column was launched in September 2013 as an educational resource to authors and to provide practical suggestions about best practices in statistical analysis and reporting.
This month we launch a new column "Points of Significance" devoted to statistics, a topic of profound importance for biological research, but one that often doesn’t receive the attention it deserves.
The "aura of exactitude" that often surrounds statistics is one of the main notions that the Points of Significance column will attempt to dispel, while providing useful pointers on using and evaluating statistical measures.
—Dan Evanko, Let's Give Statistics the Attention it Deserves in Biological Research
The column is co-authored with Naomi Altman (Pennsylvania State University). Paul Blainey (Broad) is a contributing co-author.
Free Access
In February 2015, Nature Methods announced that the entire Points of Significance collection will be free.
When Nature Methods launched the Points of Significance column over a year ago we were hopeful that those biologists with a limited background in statistics, or who just needed a refresher, would find it accessible and useful for helping them improve the statistical rigor of their research. We have since received comments from researchers and educators in fields ranging from biology to meteorology who say they read the column regularly and use it in their courses. Hearing that the column has had a wider impact than we anticipated has been very encouraging and we hope the column continues for quite some time.
—Dan Evanko, Points of Significance now free access
Also, in a recent post on the ofschemesandmemes blog, a new statistics collection for biologists was announced.
The pieces range from comments, to advice on very specific experimental approaches, to the entire collection of the Points of Significance columns that address basic concepts in statistics in an experimental biology context. These columns, originally published in Nature Methods thanks to Martin Krzywinski and guest editor Naomi Altman, have already proven very popular with readers and teachers. Finally, the collection presents a web tool to create box plots among other resources.
—Veronique Kiermer, Statistics for biologists—A free Nature Collection
continuity and consistency
Each column is written with continuity and consistency in mind. Our goal is to never rely on concepts that we have not previously discussed. We do not assume previous statistical knowledge—only basic math. Concepts are illustrated using practical examples that embody the ideas without extraneous complicated details. All of the figures are designed with the same approach—as simple and self-contained as possible.
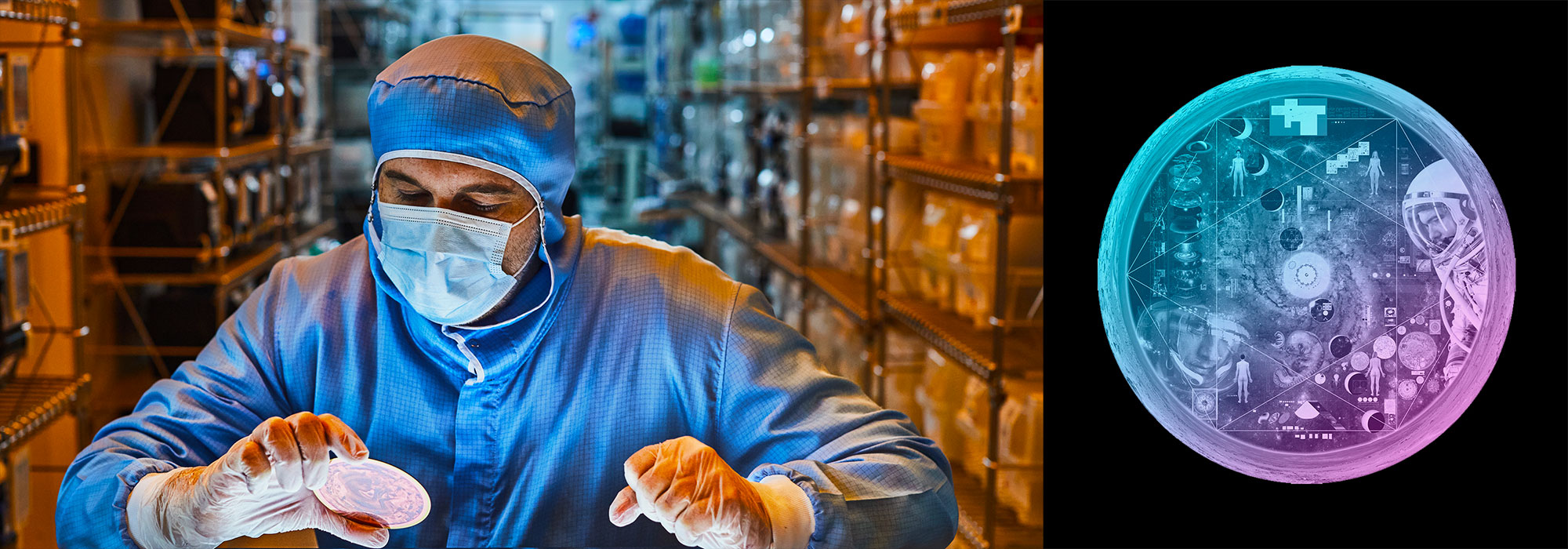
Nasa to send our human genome discs to the Moon
We'd like to say a ‘cosmic hello’: mathematics, culture, palaeontology, art and science, and ... human genomes.



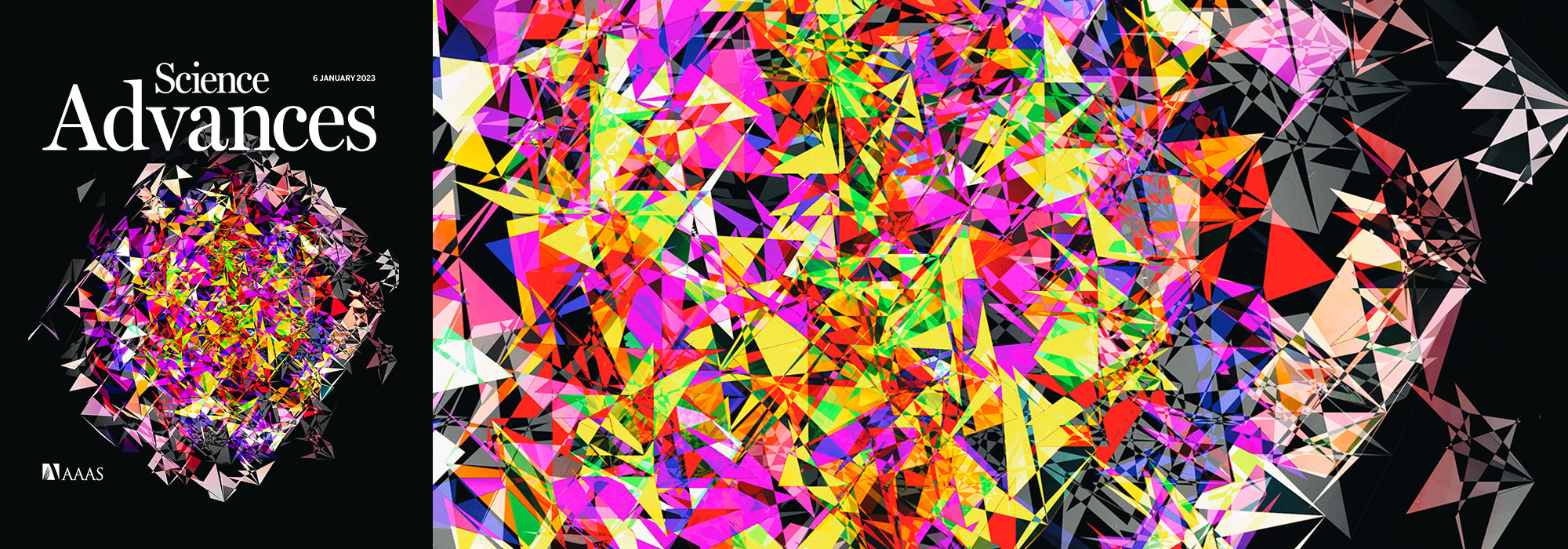
Comparing classifier performance with baselines
All animals are equal, but some animals are more equal than others. —George Orwell
This month, we will illustrate the importance of establishing a baseline performance level.
Baselines are typically generated independently for each dataset using very simple models. Their role is to set the minimum level of acceptable performance and help with comparing relative improvements in performance of other models.

Unfortunately, baselines are often overlooked and, in the presence of a class imbalance5, must be established with care.
Megahed, F.M, Chen, Y-J., Jones-Farmer, A., Rigdon, S.E., Krzywinski, M. & Altman, N. (2024) Points of significance: Comparing classifier performance with baselines. Nat. Methods 20.
Happy 2024 π Day—
sunflowers ho!
Celebrate π Day (March 14th) and dig into the digit garden. Let's grow something.

How Analyzing Cosmic Nothing Might Explain Everything
Huge empty areas of the universe called voids could help solve the greatest mysteries in the cosmos.
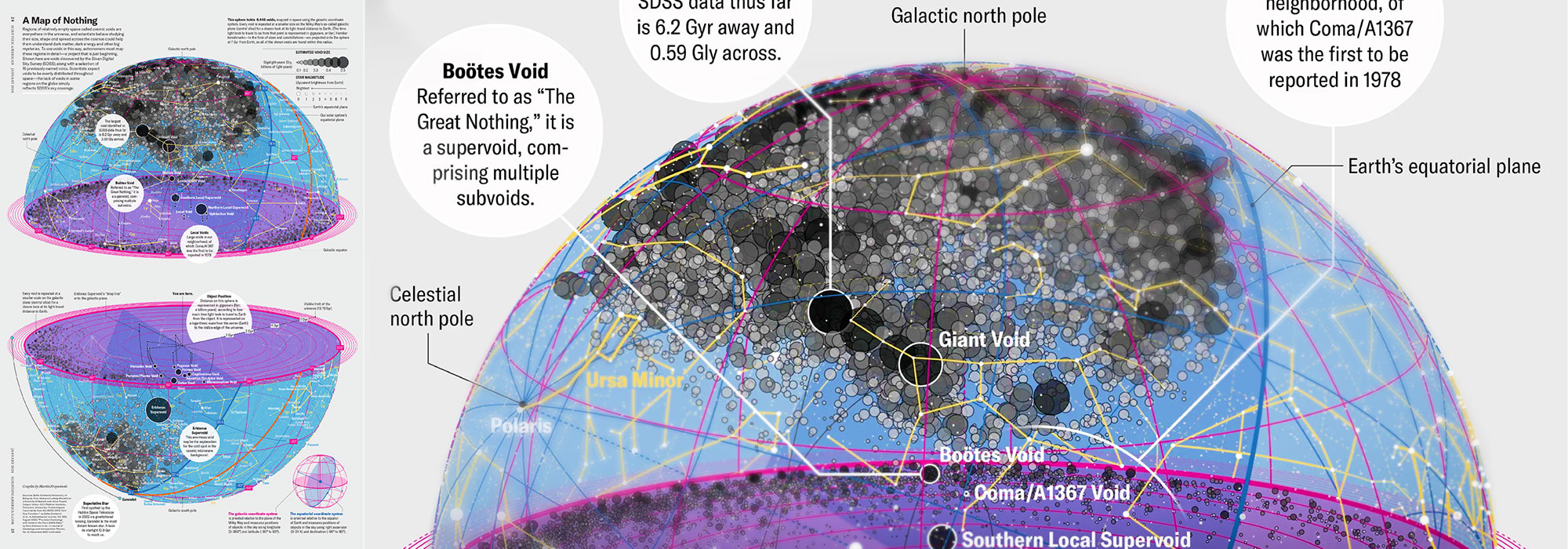
My graphic accompanying How Analyzing Cosmic Nothing Might Explain Everything in the January 2024 issue of Scientific American depicts the entire Universe in a two-page spread — full of nothing.
The graphic uses the latest data from SDSS 12 and is an update to my Superclusters and Voids poster.
Michael Lemonick (editor) explains on the graphic:
“Regions of relatively empty space called cosmic voids are everywhere in the universe, and scientists believe studying their size, shape and spread across the cosmos could help them understand dark matter, dark energy and other big mysteries.
To use voids in this way, astronomers must map these regions in detail—a project that is just beginning.
Shown here are voids discovered by the Sloan Digital Sky Survey (SDSS), along with a selection of 16 previously named voids. Scientists expect voids to be evenly distributed throughout space—the lack of voids in some regions on the globe simply reflects SDSS’s sky coverage.”
voids
Sofia Contarini, Alice Pisani, Nico Hamaus, Federico Marulli Lauro Moscardini & Marco Baldi (2023) Cosmological Constraints from the BOSS DR12 Void Size Function Astrophysical Journal 953:46.
Nico Hamaus, Alice Pisani, Jin-Ah Choi, Guilhem Lavaux, Benjamin D. Wandelt & Jochen Weller (2020) Journal of Cosmology and Astroparticle Physics 2020:023.
Sloan Digital Sky Survey Data Release 12
Alan MacRobert (Sky & Telescope), Paulina Rowicka/Martin Krzywinski (revisions & Microscopium)
Hoffleit & Warren Jr. (1991) The Bright Star Catalog, 5th Revised Edition (Preliminary Version).
H0 = 67.4 km/(Mpc·s), Ωm = 0.315, Ωv = 0.685. Planck collaboration Planck 2018 results. VI. Cosmological parameters (2018).
constellation figures
stars
cosmology